Linkage equilibrium, disequilibrium + GWAS 🙈🤷🏽♀️
Linkage disequilibrium can manage to tell us a lot or deeply confuse us about GWAS studies
Basically, for humanity to be able to reproduce, Meiosis has to occur at the individual level. Meiosis is a type of cell division in sexually reproducing organisms. During meiosis, recombination occurs—recombination is when homologous chromosomes (the same chromosome from each parent) pair up and exchange DNA.
At the scale of populations, this can lead to different rates of certain alleles, or alternative forms of a gene that can arise via mutation. It’s notable that a SNP(Single Nucleotide Polymorphism) is what creates the definition for an allele of a gene—it introduces variance. A linkage means that two alleles are located on the same chromosome.
Since recombination occurs naturally and at scale, there are two population level terms worth knowing—
Linkage equilibrium: When two or more alleles on the same chromosome occur randomly in a population. This usually means that the alleles are far enough apart on the chromosome that they are equally likely to be inherited in recombination
Linkage disequilibrium—When two or more alleles on the same chromosome to be transmitted together more or less often than expected by chance alone. This usually occurs when alleles are very close to each other on the chromosome
At the end of meiosis we have sperm or egg cells with recombinated DNA. When a sperm cell meets an egg cell during reproduction, the chromatids from each meet to form new chromosomes. This results in an embryo with a whole new genome.
What’s cool is, at the scale of the general population and seen generationally, the crossing over process is not completely random AKA the reason linkage disequilibrium is a thing.
Cheat sheet:
The higher the likelihood of two alleles staying together, the higher the linkage disequilibrium. This can be seen on in linkage disequilibrium heat maps!
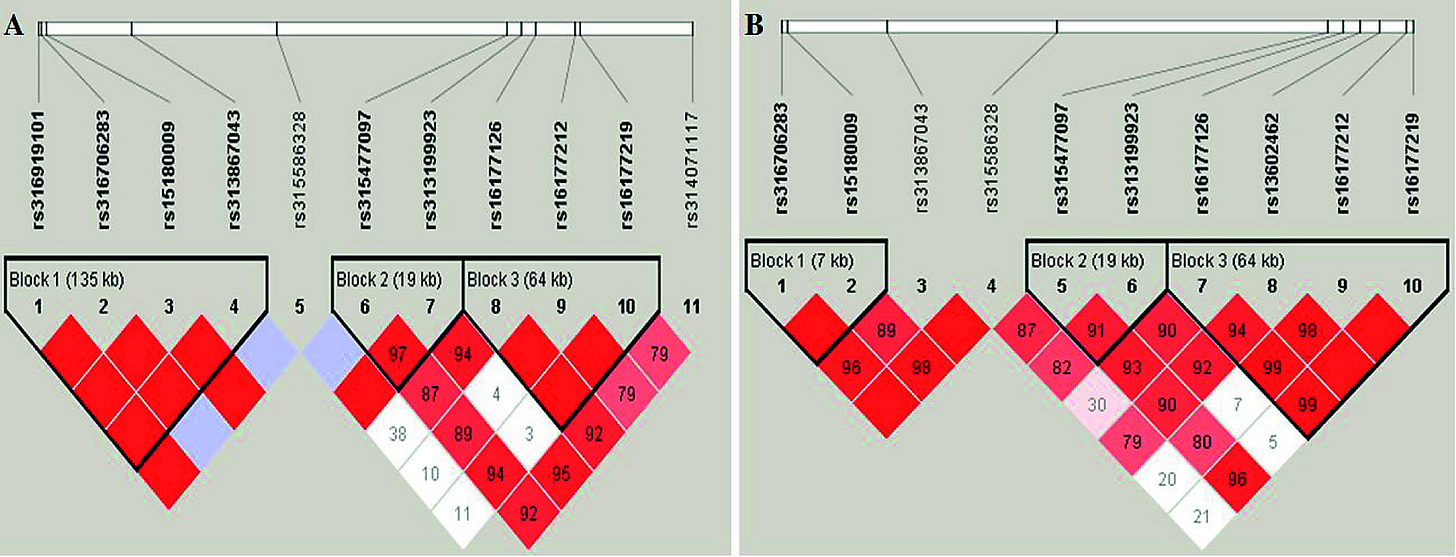
Interpreting the LD heat map below
The black triangles around red blocks represent sections of DNA that are unlikely to be separated from each other in crossing over and therefore in linkage disequilibrium with each other and are called haplotype blocks
Haplotype == A grouping of genomic variants that tends to be inherited together
Okay—parts of our genome are conserved, parts of it are less conserved… so what?
When we conduct genotyping analyses to understand which alleles are associated with which traits, linkage disequilibrium can get in the way. Occasionally multiple SNPs in one concentrated area may be identified, making it tough to account for linkage disequilibrium in Genome Wide Association Studies. New methods are on the up an up such as novel penalized regression models. It will be really interesting to see which approaches are developed to address this in the coming years!