So, what are SNPs? + Why do they matter?
Urban Dictionary defines SNP as "Smile Nose Puff." It's what most people do when they read something funny online. Feel free to smile + nose puff as you read.
In late December in 1999, the Japanese government drew together an outline of the Millennium Genome Project (MGP). The Millennium Project, at large, was an effort to get industry, academia, and government to cooperate on a wide range of R&D projects to revitalize the Japanese economy, one with an increasing elderly population. Five diseases were chosen as the targets of the project: Alzheimer's disease, cancer, diabetes, hypertension and asthma. The approach to studying these diseases included identifying candidate genes, a genome-wide + gene-based SNP scan, and statistical methods.
I will spend this article diving into what SNPs are, relevance in research, and gushing about my fascination with SNPs.
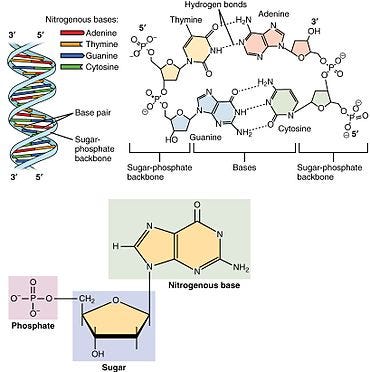
Before we dive into SNPs, review the structure of DNA nucleotides, DNA structure, and bonding using the image below (taken from Wikipedia) —
In genetics, SNP = Single Nucleotide polymorphism (pronounced snip). is a variation at a single position in a DNA sequence among individuals. Refer to the diagram above and imagine that where there is a thymine, If more than 1% of a population carries a different nucleotide at a specific position in the DNA sequence. This notable difference is classified as a SNP. In the cases where a SNP occurs in a gene, the gene is considered to have more than one allele. SNPs can also be found outside of genes— they often also occur in noncoding regions of DNA that regulate expression. SNPs are the MOST common type of DNA sequence variation there is. It is important to know that SNPs and disease causing (point) mutations are not the same! No disease causing mutation is as common as a SNP.
There are 2 types of SNPs: Linked + Causative…
Linked (also called indicative) SNPs: These reside outside genes and have no affect protein function. They do correspond to a specific drug response or to the risk for getting a certain disease.
Causative SNPs: These affect the way a protein might function, correlate with a disease, and can influence a person's response to medication. There are 2 types.
Coding —> Alter Amino Acid in protein
Non-coding —> Changes amount of a protein that is produced
SNPs can be used to functionally study a disease or at a larger scale in Genome Wide Association Studies (GWAS). My fascination is largely with how SNPs are used in GWAS.
GWAS look to identify the single nucleotide polymorphisms that are common to the human genome and look at how these polymorphisms are distributed across different populations. These GWAS studies help scientists find associations between individual SNPs and disorders that are passed on generationally, in Mendelian fashion.